Standard Python Tools¶
There are two standard “core” Python modules that provide much of the
functionality necessary to access internet resources from inside python:
urllib2 and urllib. You have seen urllib2 in each of the prior
workshops when downloading example files or datasets, like this:
import urllib2
url = 'http://python4astronomers.github.com/_downloads/image_sources.csv'
open('image_sources.csv', 'wb').write(urllib2.urlopen(url).read())
Webpage Retrieval¶
It is useful to write the example above a bit differently since readability counts and we want to highlight a few steps.
with open('image_sources.csv','wb') as outfile:
handler = urllib2.urlopen(url)
data = handler.read()
outfile.write(data)
handler.close()
First, urllib2 has returned a file like object handler that
points to a specific web resource. Various attributes about the
webpage are available:
print(handler.code)
print(handler.headers['content-type'])
In this particular example the data request is handled after the
initial response has been examined; the data is streamed into a local
variable data and then saved to a local file. As demonstrated in
the ASCII files portion of the Reading
and Writing Files workshop, one alternatively could
parse the returned data stream into a table for further use.
Building Queries¶
Now we are getting closer to searching for data online via python. Traditionally, each data archive has implemented its own set of query parameters; more often then not you experience it as a web form with lots of check and input boxes. Most of the time these web forms are simply input wrappers that build a query string and process the result.
Here is a very simple query example: search for HST images near M 51. It is worth noting that you have to find and read the MAST HST documentation to learn which parameters are needed to build the query.
import urllib2, urllib
# this is the query url for this service.
url = 'http://archive.stsci.edu/hst/search.php'
# make a dictionary of search parameters
p = {}
p['target']="M 51"
p['radius']=1.0
p['max_records']=10
p['outputformat']='CSV'
p['action']='Search'
# encode the http string
query = urllib.urlencode(p)
# create the full URL for a GET query
get_url = url + "?" + query
# create the GET handler
handler = urllib2.urlopen(get_url)
# check it
print(handler.code)
print(handler.headers['content-type'])
# save it
with open('hst_m51.csv','wb') as f:
f.write(handler.read())
The magic is that urllib module takes care of encoding the parameters using standard HTTP rules. You can compare the input dictionary key,value pairs with the HTTP url encoding.
# some simple formatting where the format %-M.Ns means
# "-" :: 'left justified'
# "M" :: 'minimum string length, padded'
# "N" :: 'maximum string length, sequence sliced'
# "s" :: 'format as string'
sform = "%-20.20s %-10.10s %-30.30s"
# make a header
print(sform % ('parameter','value','encoded'))
print(sform % (3*(100*'-',)))
# for each paramter show the input items from the dictionary
# "p" and the output query string.
for a,b in zip(p.items(),query.split('&')):
print(sform % (a+(b,)))
Exercise: import WISE catalog data for a young cluster
In this exerice you will use a different service (IRSA) and with a different overall goal for these data.
import atpy, urllib, urllib2
url = "http://irsa.ipac.caltech.edu/cgi-bin/Gator/nph-query"
p = {}
p['spatial'] = "Cone"
p['objstr'] = "IC 348"
p['outfmt'] = 1 # IPAC table format
p['catalog'] = 'wise_prelim_p3as_psd'
p['radius'] = 300
query = urllib.urlencode(p)
get_url = url + "?" + query
handler = urllib2.urlopen(get_url)
raw = handler.read()
print(raw[0:255])
with open('ic348_wise.tbl', 'wb') as f:
f.write(raw)
The challenge is to immediately analyze the results of this query. The exercise is to make a simple color-color plot of these objects.
Click to Show/Hide Solution
There are MANY ways we have looked at in the workshops for converting this result to a numpy array. Some of these examples parse the raw web data directly, circumventing a need to write it to file. Some use different means to try to deal with the data types and null values in the result.
t1 = atpy.Table('ic348_wise.tbl',type="ipac")
t2 = asciitable.read(raw, Reader=asciitable.IpacReader)
t3 = atpy.Table(raw,type='ascii')
fill_values = [('null',numpy.nan)]
t4 = asciitable.read(raw, Reader=asciitable.IpacReader, \
fill_values=fill_values)
t5 = atpy.Table(raw,type='ascii', fill_values=fill_values)
Its important to realize that YMMV as to how these differ in their output. For example, lets look at the output table types and the data types for a couple of columns:
t = [t1, t2, t3, t4, t5]
for i in t:
c = (type(i), i['j_m_2mass'].dtype, i['tmass_key'].dtype)
print("%40s%10s%10s" % c)
The output table types (and hence their built-in utilities), column data types
and treatment of null values vary and this matters when trying to make a plot
or perform other types of analysis. We will use t1 as the data types are
correct. It also preserves more of the metadata of the query.
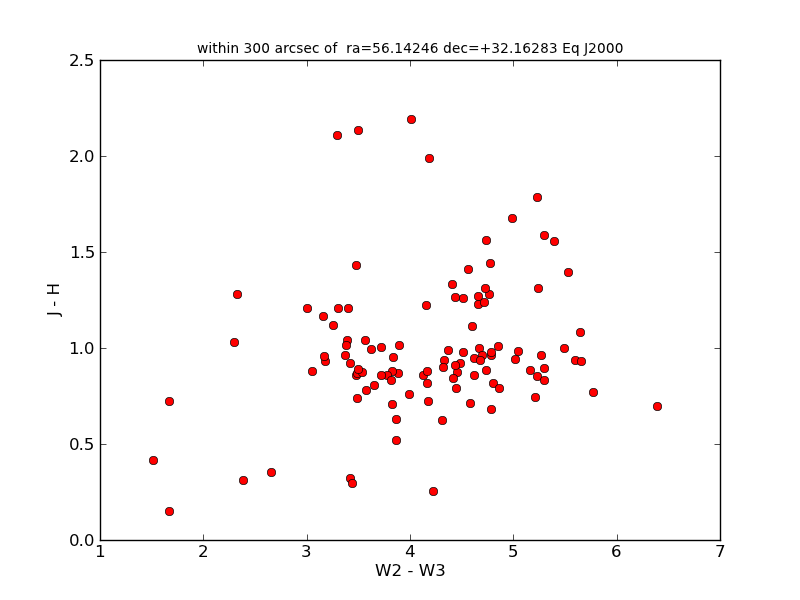
Just a quick plot that reuses some of this metadata in the plot title.
clf()
plot(t1['w2mag']-t1['w3mag'], t1['j_m_2mass']-t1['h_m_2mass'], 'ro')
xlabel('W2 - W3')
ylabel('J - H')
title(t1.keywords['SKYAREA'], fontsize='small')
Read the instructions!
Because these web interfaces vary from archive to archive it is worth emphasizing that building the query string for a particular data archive begins with reading the documentation page for that archive’s GET interface.
Here are some links to these documentation pages for some archives
and the service url for these are: